New research by Sales and colleagues looks at the monitoring of terrestrial mammal communities and compares the efficacy of landscape-level monitoring using environmental DNA (eDNA) to that of conventional methods. Here the authors summarise their findings.
Accurately and effectively monitoring biodiversity is a key consideration in this rapidly changing world. Consistent and regular monitoring of species communities is pivotal for ongoing management, conservation and policy implementation within ecosystems. Over the last decade, the use of environmental DNA (eDNA) as a method to detect and monitor individual species or whole communities (eDNA metabarcoding) has drastically increased. eDNA is DNA left in the environment by organisms, often in the form of faeces/urine, blood, hair or flakes of skin; and it can be recovered from water, soil or even air.
To date, most applications of eDNA have been focused on monitoring aquatic organisms such fishes or macroinvertebrates. Mammals, among which are some of the most imperiled taxonomic groups in terms of threatened or endangered status, have remained relatively neglected in eDNA studies up until recently. . By comparing eDNA metabarcoding against traditional latrine (faecal) surveys and camera trap monitoring, we were able to explicitly show that the general mammal community and specific target mammals, both semi-aquatic and terrestrial, could be reliably detected using eDNA-based survey methods.
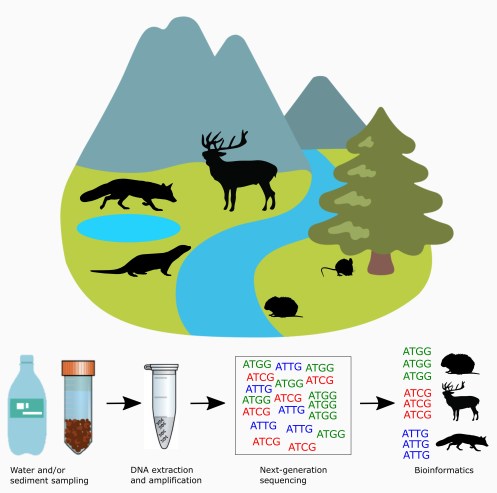
Landscape-scale monitoring
Using occupancy/detection analysis, we tested the ability of each of the three survey methods (eDNA metabarcoding, latrine and camera traps) to detect multiple mammal species in multiple sites in Scotland, UK. We focused on three species primarily: water voles (a threatened semi-aquatic species), field voles (a common, terrestrial small mammal), and red deer (a common, wide-ranging terrestrial species). Although these species were explicitly targeted, we were also able to recover the majority of the expected mammal community across study sites, with 23 wild mammals detected. Water-based eDNA methods were able to detect the expected mammal communities based on historical data and sometimes provided higher resolution information on species than camera traps (which often have to be deployed to maximise either field-of-view or focal length depending on the suite of species of interest).

Comparing surveying methods
All methods are not created equal. For example, to get a 95% chance of detecting the water vole, you would need three latrine surveys, four water-based eDNA samples or five weeks of camera trapping a sampling location. A consideration of this ‘effort’ of each method is important. Take the field vole as example, where it would take upwards of 14 weeks of camera trapping to achieve the same detection results as only six water-based eDNA samples or five latrine surveys; the cost and time differential between collecting water samples from specific points versus five repeat cross-country treks to survey entire habitat patches is immense.
However, there are some downsides to using eDNA, as demonstrated in this study. Although not explicitly analysed for detection, eDNA failed to detect wide-ranging predators such as weasels and otters, even when camera traps regularly detected these species. As such, different methods, each with its own benefits, could be used in combination to maximise monitoring protocols depending on the species of interest from a management or conservation perspective.
Application of eDNA to monitor mammals
The application of eDNA metabarcoding is a relatively new technique in biomonitoring and therefore isn’t without its drawbacks. However, based on the results of this study, and other recent studies in other geographic regions, the use of eDNA is a helpful addition to the manager’s tool kit for monitoring the terrestrial and semi-aquatic mammal community. It is an even more powerful method when used to compliment other non-invasive surveying methods. Consider that an entire watershed could be monitored from carefully selected sampling locations along a riparian corridor. This will pick up not just the semi-aquatic species in regular contact with the water, but most, if not all, the terrestrial species also. These samples could be used to identify cryptic, threatened, or invasive species; then more intensive and spatio-temporally precise methods can be employed to get a higher resolution of presence/absence.
The cost of biomonitoring with eDNA is continuing to decline, and in the foreseeable future, this may be a reasonable way to accumulate a local ‘library’ of community or biodiversity data based on strategic eDNA sampling on regional or national levels. This knowledge would let managers deploy more costly, time-intensive, or invasive methods like trapping more tactically. The entire community and specific species could be targeted with the use of eDNA, a method that seems to perform just as well, if not better, than other non-invasive survey methods for certain groups within mammals (e.g. group-living species).
Read the full article, Fishing for mammals: Landscape‐level monitoring of terrestrial and semi‐aquatic communities using eDNA from riverine systems, in Journal of Applied Ecology.

Reblogged this on The Secret Life of a Field Biologist and commented:
Check out this research blog, that I just happened to write. It’s on a new paper that came from a large, international collaboration.
LikeLiked by 1 person